Introduction
microRNAs (miRNAs) are ~22 nucleotide (nt)-long single stranded RNA molecules that form a complex with the Argonaute protein to bind primarily to the 3’UTR of target messenger RNAs (mRNAs) which leads to the repression in gene expression. It is widely accepted that miRNAs act as fine-tuners of gene expression and are reported to regulate at least 60% of the mRNAs. They are involved in various biological processes with implications in many diseases such as cancer. The miRNAs bind to their target mRNAs through Watson-Crick nucleotide matches in which the pairings at the 6-nt seed region have been shown to play an important role. Previous empirical computational analyses have identified that 6-nt perfect WCPs are the primary functional interactions between miRNAs and their mRNA targets, and the four site types have been regarded as the canonical site types (CSTs). However, there exist an enormous amount of possible site types between the miRNAs and human mRNA targets in which only a small fraction has been evaluated by previous studies.
microRNAs (miRNAs) are ~22 nucleotide (nt)-long single stranded RNA molecules that form a complex with the Argonaute protein to bind primarily to the 3’UTR of target messenger RNAs (mRNAs) which leads to the repression in gene expression. It is widely accepted that miRNAs act as fine-tuners of gene expression and are reported to regulate at least 60% of the mRNAs. They are involved in various biological processes with implications in many diseases such as cancer. The miRNAs bind to their target mRNAs through Watson-Crick nucleotide matches in which the pairings at the 6-nt seed region have been shown to play an important role. Previous empirical computational analyses have identified that 6-nt perfect WCPs are the primary functional interactions between miRNAs and their mRNA targets, and the four site types have been regarded as the canonical site types (CSTs). However, there exist an enormous amount of possible site types between the miRNAs and human mRNA targets in which only a small fraction has been evaluated by previous studies.
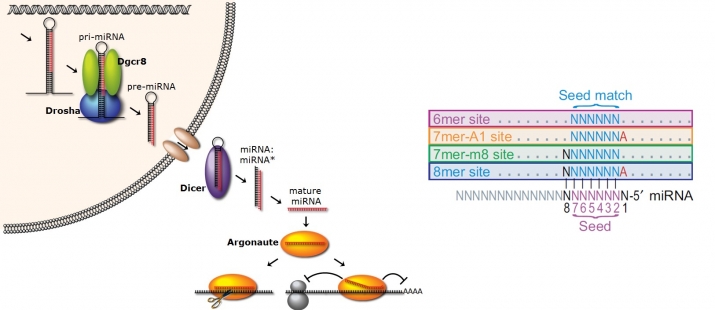
(miRNA function & canonical targeting rules (right, Baek et al. 2008))
Pipeline for discovery of functional miRNA targeting rules
In order to overcome the limited number of evaluated site types from previous studies, our lab systematically determined all of the interactions that can possibly occur between the miRNA and target mRNAs by utilizing large-scale microarray data measuring the transcriptome response to ectopically introduced miRNAs combined with the development of a powerful analysis pipeline. In doing so, we statistically evaluated whether each of the >2 billion unique miRNA-mRNA interactions were enriched in highly downregulated genes in response to overexpressed miRNAs.
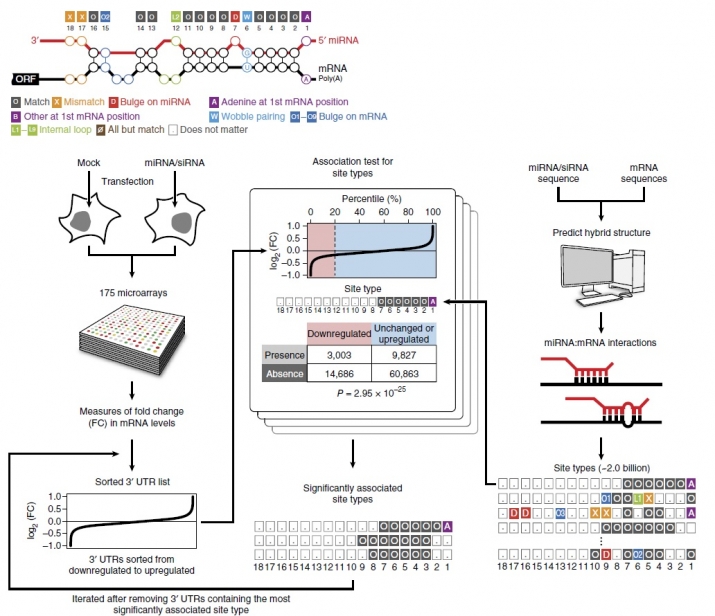
(miRNA targeting analysis pipeline (Kim, Sung, Park et al. 2016))
General rules for functional miRNA targeting
Through our massive-scale study, seven non-canonical site types (NSTs) were detected in addition to the four previously known CSTs. These NSTs were validated using independent microarray data as well as through experimental assays. Importantly, most of the NSTs were found to exhibit significant conservation across vertebrate genomes indicating that they likely play biologically relevant roles. Although the level of repression elicited by these NSTs on the transcriptome is much weaker than that of the CSTs, the large number of mRNAs targeted by NSTs suggest that their overall impact on the transcriptome is comparable to that of CSTs. Our unprecedented study in determining the comprehensive rules for functional miRNA targeting identified novel NSTs, expanding not only the functional targeting repertoire of the miRNAs but also revealing the complex and intricate nature by which the miRNAs impact the gene regulatory network (Kim, Sung, Park et al. 2016). These findings from our comprehensive study help us better understand the mechanisms of miRNA-mediated gene regulation, and our lab will further conduct research relevant to this study.
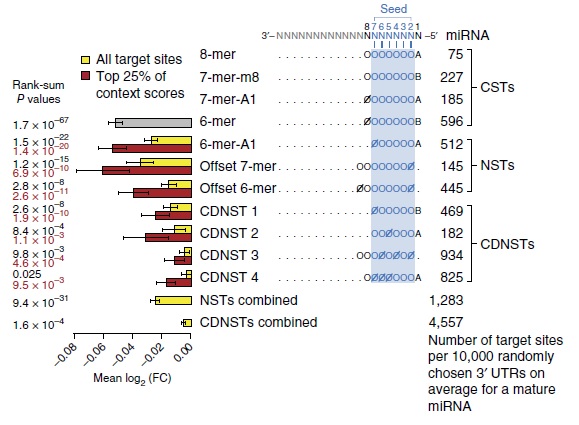
(Discovery of functional miRNA targeting rules (Kim, Sung, Park et al. 2016))
References
1. Baek, D., Villén, J., Shin, C., Camargo, F. D., Gygi, S. P., & Bartel, D. P. The impact of microRNAs on protein output. Nature (2008).
2. Kim, D., Sung, Y. M., Park, J., Kim, S., Kim, J., Park, J., ... & Baek, D. General rules for functional microRNA targeting. Nature Genetics (2016).
